A new study by scientists at Kolkata’s Bose Institute has challenged a long-accepted model of how bacteria regulate gene expression, a finding that could help open fresh avenues in the fight against tuberculosis and other bacterial infections. The research suggests that a mechanism long treated as universal in molecular biology does not apply uniformly to Mycobacterium tuberculosis, the bacterium that causes TB.
For decades, scientists believed that a protein known as the sigma factor binds to RNA polymerase to initiate bacterial transcription and then detaches once RNA synthesis moves into the elongation phase. This “sigma-cycle” had widely been considered a standard feature of bacterial gene regulation, including in TB bacteria. But the new study shows that the process is more complex and far less uniform than previously understood.
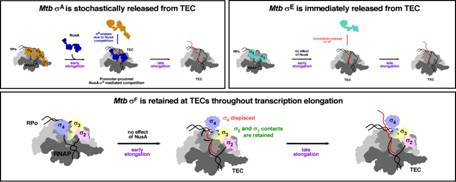
The work, carried out by Dr. Jayanta Mukhopadhyay and Dr. N. Hazra, found that different sigma factors in M. tuberculosis behave differently during transcription. While sigma A and sigma E detach from RNA polymerase either immediately or progressively during elongation, sigma F remains stably attached even as transcription continues. The findings were published in Nucleic Acids Research.
That distinction is important because sigma F is associated with stress survival and adaptation in TB bacteria. Its continued attachment to RNA polymerase suggests a previously unrecognised way by which the pathogen may sustain expression of stress-response genes inside the host, potentially helping it survive under hostile conditions.
Researchers used a combination of in vitro transcription assays, fluorescence-based measurements, high-resolution protein interaction studies, and in vivo validation through chromatin immunoprecipitation followed by quantitative PCR to arrive at the finding. The study argues that these variable sigma–RNA polymerase interactions could offer highly specific targets for future antimicrobial development. Instead of targeting enzyme active sites, where drug resistance often emerges, future therapies may be able to disrupt critical protein–protein interactions essential for bacterial survival.
The findings matter in the broader context of rising antimicrobial resistance and the continuing global burden of tuberculosis. By revising a textbook understanding of bacterial transcription, the study adds a basic-science insight that may eventually support the development of more targeted anti-TB strategies.
Reference: PIB
You may also like
-
Vikram-1 Heads to Sriharikota as India Moves Closer to Its First Private Orbital Rocket Launch
-
India Launches Indigenous Silicon Photonics Design and Testing Platforms in Chennai
-
RRI Scientists Discover Two New Flow Patterns In Clay, Opening Fresh Insights For Oil Recovery And Material Transport
-
National Institute of Technology Rourkela Develops Patented Bio-Ink for Bone and Cartilage Repair
-
India Identified as Key Centre in Jamun’s Evolutionary History