In a breakthrough that challenges nearly five decades of biological understanding, scientists from the Bose Institute and Rutgers University have overturned a central textbook model of how bacteria regulate their genes. The findings, published in the prestigious journal Proceedings of the National Academy of Sciences, could reshape modern microbiology and open new pathways for antibiotic development and biotechnology innovation.
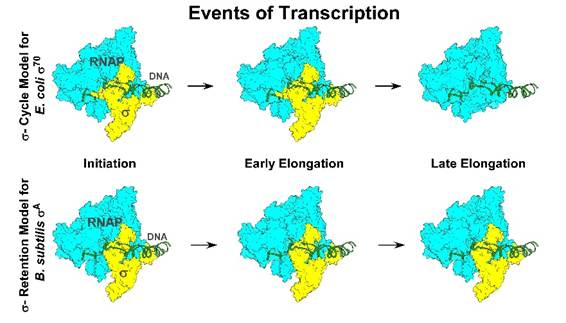
For almost 50 years, biology textbooks have described bacterial gene activation through what is known as the “sigma (σ) cycle.” According to this long-standing model, sigma factors — proteins that bind to RNA polymerase — initiate transcription by helping the enzyme attach to DNA. Once transcription begins, these sigma factors were believed to detach, allowing RNA polymerase to continue copying genetic information.
What Is Transcription?
Transcription is the process by which a cell copies information from DNA into RNA.
Think of DNA as a master instruction manual. RNA polymerase is the machine that reads those instructions and creates a working copy (RNA), which then helps produce proteins — the building blocks of life.Sigma factors are helper proteins that guide RNA polymerase to the correct starting point on DNA.
This understanding was largely based on studies of the bacterium Escherichia coli, particularly its primary sigma factor known as σ70. Because E. coli has been one of the most widely studied bacteria in the world, its mechanisms were assumed to represent a universal rule across bacterial species.
However, the new research demonstrates that this model does not apply to all bacteria.
The team discovered that in Bacillus subtilis, the principal transcription initiation factor σA does not detach after initiation. Instead, it remains bound to RNA polymerase throughout the entire transcription process. In addition, a modified version of E. coli’s σ70 — lacking a structural region called domain 1.1 — behaved similarly and remained attached.
“Our work shows that in Bacillus subtilis, the σA factor stays attached to RNA polymerase all the way through the transcription process,” said Dr. Jayanta Mukhopadhyay of Bose Institute, the corresponding author of the study. “This fundamentally changes how we think about bacterial transcription and gene regulation.”
Why Is This a Big Deal?
For decades, scientists believed sigma factors always followed a “bind-and-release” cycle.
This study shows that in some bacteria, sigma factors do not release.
That means the long-accepted “sigma cycle” model is not universal.In science, overturning a 50-year-old model is rare — especially one taught globally in textbooks.
Using advanced methods such as biochemical assays, chromatin immunoprecipitation, and fluorescence-based imaging, researchers observed sigma factor behaviour in real time. These cutting-edge tools allowed them to track protein interactions inside living systems with remarkable precision.
Their experiments revealed that Bacillus subtilis σA and the altered E. coli σ70 variant remain stably associated with transcription complexes. By contrast, the full-length E. coli σ70 detaches randomly, or stochastically, during elongation.
“These findings provide compelling evidence that the long-accepted σ cycle does not apply to all bacteria,” noted co-author Aniruddha Tewari of Bose Institute. “It opens new avenues for understanding bacterial gene regulation and its evolution.”
How Could This Help Medicine?
Many antibiotics work by targeting bacterial gene expression.
If scientists now understand transcription more accurately, they can:• Design better antibiotics
• Develop molecules that block infection mechanisms
• Create precision drugs targeting bacterial regulation systems
This is especially important as antimicrobial resistance continues to rise globally.
The implications extend beyond medicine. A revised understanding of transcription mechanisms could influence research in bacterial physiology and stress response. It could also help scientists engineer microorganisms more efficiently for industrial applications.
Beyond Antibiotics — Industrial Impact
Better control of bacterial gene regulation can help design microbes that:
• Produce biofuels more efficiently
• Manufacture biodegradable plastics
• Generate therapeutic compounds
• Improve fermentation technologies
In simple terms, mastering gene regulation means mastering microbial productivity.
The study, authored by Aniruddha Tewary, Shreya Sengupta, Soumya Mukherjee, Nilanjana Hazra, Yon W. Ebright, Richard H. Ebright, and Jayanta Mukhopadhyay, represents a significant India–US scientific collaboration.
By challenging a biological rule that has stood for nearly half a century, Indian scientists have not only rewritten a chapter in molecular biology but also expanded the frontier of genetic research. In doing so, they have demonstrated how fundamental science can reshape medicine, biotechnology, and our understanding of life at its most microscopic level.
Publication link: doi:10.1073/pnas. 2503801122
Refernce: PIB
You may also like
-
MeitY Unveils Indigenous 30 kW WBG-Based Integrated Drive System to Power India’s EV Revolution
-
CeNS Researchers Unlock Enhanced Optical Performance with Gold–LC Composite
-
Sarvam AI: Powering a Made-in-India AI Revolution
-
Csir-ampri Designed & Developed Sodar System Facility Inaugurated At IMD
-
India Will Soon Come Up With E-observer System For Remote Monitoring Of Fishing, Says Union Fisheries Secretary